A molecule of RNA consists of a long chain of subunits, called ribonucleotides. Each ribonucleotide contains one of four possible bases: adenine, guanine, cytosine, or uracil (abbreviated as A,G,C,U respectively). It is this sequence of bases, known as the primary structure of the RNA that distinguishes one RNA from another.
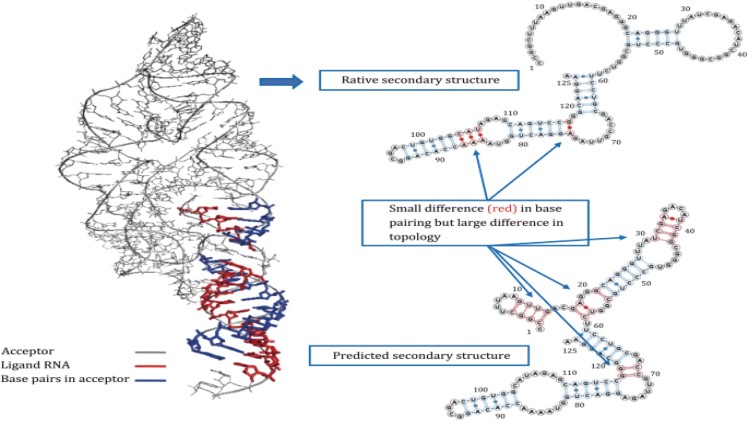
Under normal physiological conditions, a ribonucleotide chain can bend back upon itself, and the bases can hydrogen-bond with one another, such that the molecule forms a coiled and looped structure. The pattern of hydrogen bonding is generally called the secondary structure, while the conformation of the molecule in 3-dimensional space is called the tertiary structure.
buy Xenical generic rxxbuynoprescriptiononline.net over the counter
.The base-to-base interactions that form the RNA secondary structure are primarily of two kinds — hydrogen bonding between G and C and hydrogen bonding between A and U, as was first described by Watson and Crick in [1953]. (See Figure 1.) In fact, there is evidence of non-Watson-Crick base pairing in such nucleic acids as the tRNAs, but these are considered to derive from the tertiary structure produced by large regions of secondary structure containing Watson-Crick base pairing.
How can you know best online technorati website, and more segfault great website
For the sake of simplicity, such base pairing is mostly ignored in this paper. Genetic information, the set of instructions that directs cell maintenance, growth, differentiation, and proliferation, is encoded in DNA molecules. RNA serves two biological purposes: It is the means by which information flows from DNA into the production of proteins, the catalysts and building blocks of cells; it also acts as a structural component of ribosomes and other complexes.
buy clomid generic rxxbuynoprescriptiononline.net/clomid.html over the counter
It is the secondary structure, and the resulting tertiary structure, that determine how the RNA will interact and react with other cell components. Work on the determination of RNA secondary structure has been carried out for decades by a number of research groups. The classical approach is direct observation of a molecule’s secondary structure using X-ray crystallography.
Make use of free SEO tools such as the MozBar to determine how authoritative the 7starhd.la is. Guest posts can also generate more traffic to your website, so aim for these sites.
How do I submit a guest post to a website? First of all, you should check the guidelines of the website like fcstream.info where you’re submitting the guest post.
To find kamitamika.com website, start by reading the content on the site. Read the posts on the blog and see what topics are popular. If your topic is not on their blog, it’s best to look for a site that does.
In addition to footwear, Woodland also produces apparel and accessories. The brand is known for its unique features and maintains high standards in its operations. The company uses advanced materials and construction techniques, as well as socially and environmentally friendly practices. And because of their dedication to sustainability, they’ve become a popular brand for outdoor enthusiasts and active lifestyle enthusiasts in many countries around the world.
Conclusion:
More indirect methods involve specific cleavage of the RNA by enzymes called ribonucleases. Much research has gone into the promising approach of computational prediction of secondary structure from knowledge of primary structure. The general method has been to search for configurations of maximum base-pairing or of minimum free energy.
Extratorrent is very famous for downloading movies but you can easily download movies from moviespur
Read More About: sattamataka143
To know more information about SEO guest Post on instazu and picdeer. You may rank your website for sharing your blog post on amihub and cinewap. Click here: Timesweb.