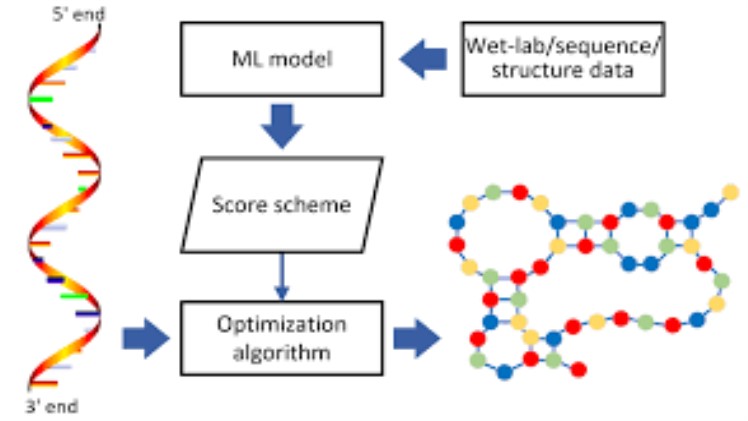
In terms of the search model discussed above, our neural network method STEEG 135 may be described as a highly parallel distributed search, wherein each possible RNA secondary structure representation is distributed over many “processing units” (one unit for each possible base-pairing) and wherein several potential secondary structures for the input sequence are represented simultaneously.
Conflict (w.r.t. constraint violation) between possible substructures is implemented by inhibitory connections between units in the respective substructures, and support (stem compatibility) is implemented by excitatory constraints.
Points in the 2˚RNA search space are considered many at a time, as they compete during the MFT network relaxation process.
buy kamagra online https://www.mabvi.org/wp-content/themes/mabvi/images/new/kamagra.html no prescription
The MFT relaxation algorithm is intended to avoid bad locally-optimal points in the space in favor of more globally-optimal solutions.
buy wellbutrin Canada langleyrx.com/wellbutrin.html no prescription
The MFT learning algorithm is intended to make this search easier by refining the parameters of this competition over many trials with a training set of sequence and structure data.
Concluding Remarks:
Connection weights constrain the dynamics of network relaxation, and can be seen as an implicit representation of knowledge, both analytic and heuristic, that aids the search process by pushing the network state transition process in particular directions and towards particular solutions in the solution space (π, {T(P)}).